Free and open source software for the HiSeq 2000/2500
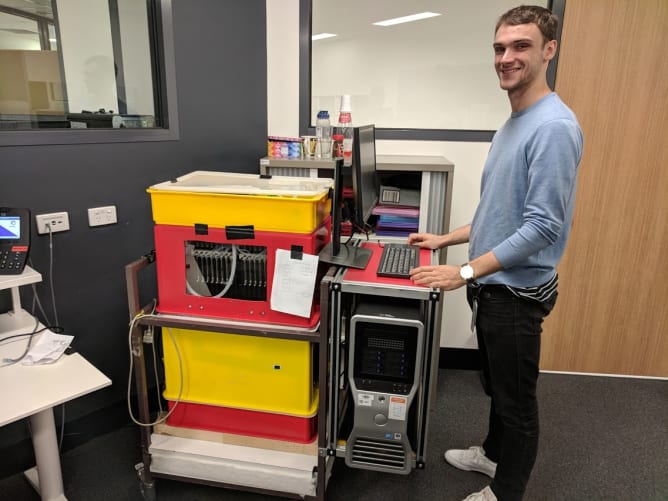
The Illumina HiSeq 2000 and 2500 are advanced DNA sequencing machines that have been superseded by a new generation that can do the same thing more quickly and cheaply. Institutes and companies around the world are now decommissioning them and they are available for a fraction of the original cost, sometimes even free.
The purpose of this campaign is to fund software development to control all the components of the HiSeq and extend its life by enabling applications other than DNA sequencing such as microscopy and novel high throughput assays. Possible uses include ongoing research, but also new citizen science projects and educational outreach.
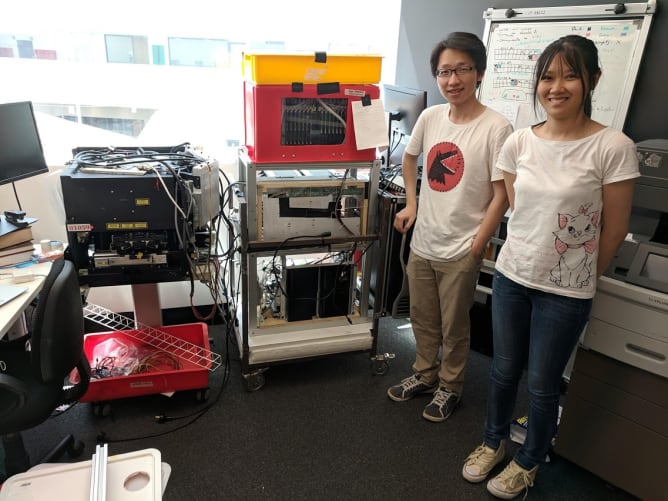
We are a community of people who met through the GOSH ( http://openhwardware.science ) and Hackteria ( https://hackteria.org ) networks. The bulk of this software development will be carried out by Kaspar Emanuel and Bengt Sjölén with others assisting where necessary.
Old Dog, New Tricks
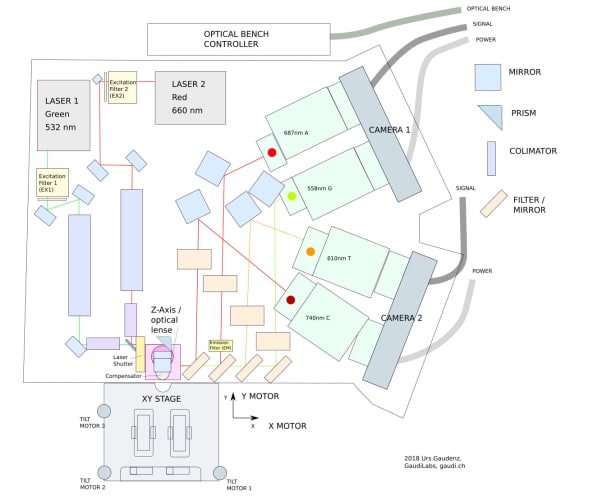
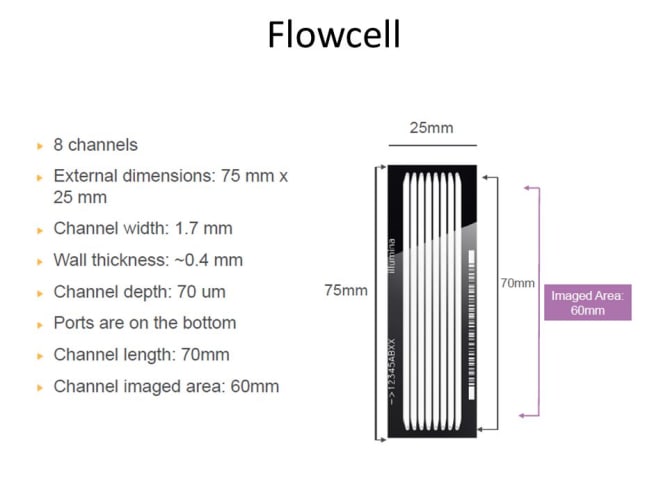
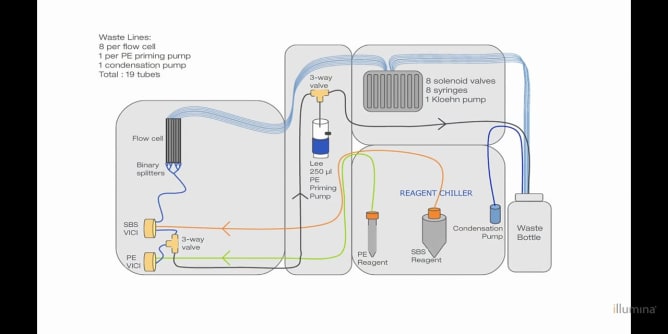
The original HiSeq 2000s cost around $650,000 in 2012 and thousands were sold around the world. These instruments contain high resolution fluorescent microscopes capable of scanning a microtiter plate-sized area. The hardware inside includes a precision stage, an optical bench supporting simultaneous four-channel laser-illuminated fluorescence acquisition, four Hamamatsu line scanning sensors, on-stage temperature control and a 16 channel microfluidic system for reagent dispensing. With open source control software it will be possible to use the old HiSeqs not only for DNA sequencing but also for various custom assays and standard fluorescent microscopy. While the ability to implement custom assays is useful for research scientists, we also hope that cheap access to a high quality fluorescent microscope will be appreciated by would-be scientists, educators and DIY biology communities.
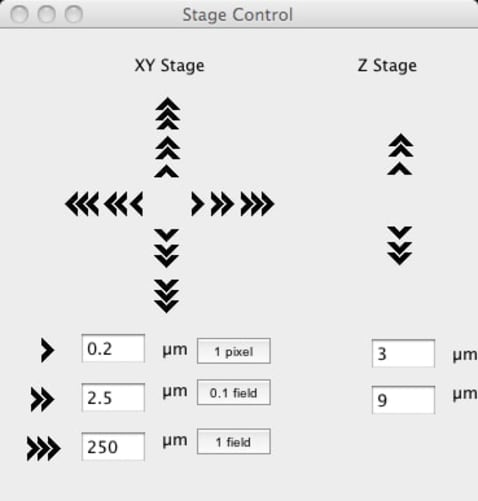
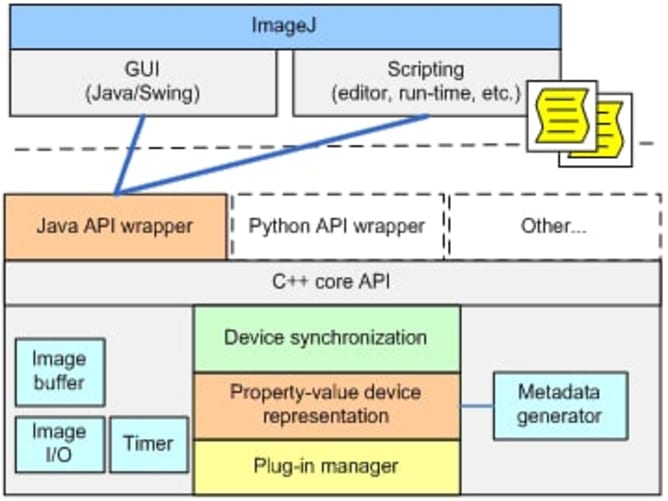
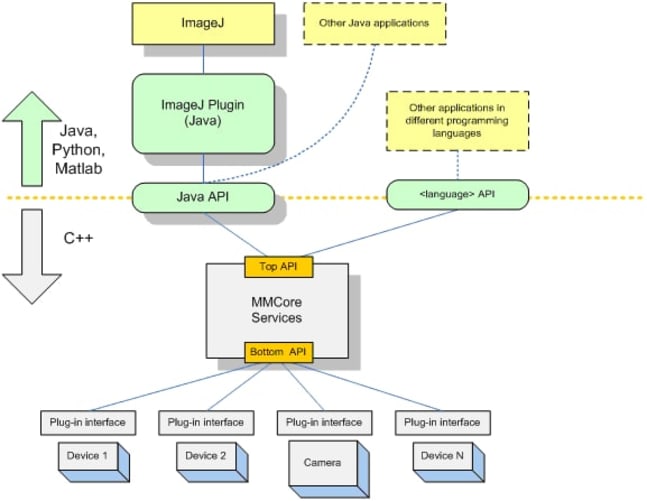
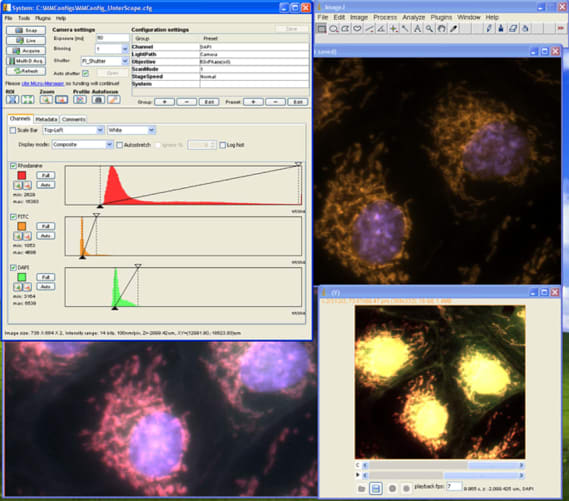
We will not be starting from scratch but intend to take a similar approach to what others have done for the Illumina GAIIx sequencers and integrate the HiSeq into a widely-used and well-supported open source microscope project, Micro-Manager. This would enable the HiSeq to immediately be usable as a fluorescent microscope via the existing Micro-Manager user interface which provides interactive controls for illumination, focus, stage positioning and image manipulation. We will write a new user interface plugin to similarly control the other HiSeq hardware such as fluid handling and temperature control. All of the functionality that we add will be available within Micro-Manager via its Java/Beanshell and Clojure scripting environments and from other applications via external calls to the core API libraries in either Python, Java, C++ and MATLAB. This means that the software components that we make can be reused by others to form the basis of whatever custom assays and novel applications they can dream up.
The Gory Details
The integration task consists of two parts. First, DeviceAdapters need to be written in C++ to enable Micro-Manager to talk to the 20+ individual components that the HiSeq contains. We have already met and spent a couple of days together establishing that we have all the information and resources needed to do this. A possible sticking point was writing the camera DeviceAdapter for the custom line scanning hardware but we now have one written and working to the point that we are confident we have it covered. The rest of the components communicate via RS-232 which has been well documented by Urs. In addition, most of the components are available separately from their suppliers along with documentation explaining their operation and command sets. Second, once the DeviceAdapters are done, a user interface needs to be written to interactively monitor and control those components not used in the microscope. This is to be done in Java using the Micro-Manager plugin framework.
The money raised will be used to feed struggling open source developers and students who are willing and able to help. If time allows, we would also like to develop and make available the design for an LED-based illumination module for transmission imaging. We will make our code available in a github repository and push updates as we go so that you can follow our progress..
This is quite a low budget and project being done mainly out of interest and on our own time so we are unable to offer any substantial inducements.
For those wanting more detail, especially about assay development, institutional involvement and other ways to contribute we encourage you to visit the ReSeq website and FAQ or to post questions to the Hackteria forum.